 
Some SR proteins can recruit the nuclear export factor 1 (NXF1) to bind RNAs, leading to the export of mRNA to the cytoplasm ( 18– 20). Consistent with their role in co-transcriptional splicing, SR proteins are present at sites of transcription and can be detected on chromatin by ChIP ( 13, 16– 17). Recent genome-wide studies have shown that SR proteins preferentially bind exonic sequences-possibly because of the higher abundance of exonic sequences in total cellular RNA-but also have a great number of binding sites in intronic regions ( 11– 15). SR proteins bind exonic and intronic splicing enhancers (ESEs and ISEs) to promote the inclusion or exclusion of exons. The serine-arginine rich splicing factors, SR proteins, are a highly conserved family of RBPs that regulate Pol II transcription, pre-mRNA splicing, polyadenylation, nuclear export, translation and stability ( 9, 10). In the cytoplasm, RBPs regulate mRNA localization, translation, stability, and degradation. Splicing continues in the nucleoplasm, where mRNP assembly for export is finalized ( 8). Historically, RNPs containing pre-mRNAs were termed heterogeneous nuclear ribonucleoprotein particles (hnRNPs), which may be expected to include both nascent RNPs and those released from chromatin by polyadenylation cleavage. Co-transcriptional RNA binding produces nascent RNPs, which lie adjacent to the DNA axis ( 7). This co-transcriptional binding is a fundamental feature in pre-mRNA maturation, which regulates co-transcriptional processing steps like capping and splicing ( 5, 6). Many RBPs bind pre-mRNAs during transcription by RNA Polymerase II (Pol II). RNP maturation is likely a dynamic process involving the binding and release of multiple factors that occurs on chromatin, within the nucleoplasm, and in the cytoplasm. The composition of RNPs thereby determines the fate and function of all RNAs ( 1). Accordingly, RNPs contain different proteins, depending on the RNA class and sequence as well as the stage of maturation. RBPs regulate and often catalyze essential steps in the processing and function of coding and non-coding RNA including: 5′ end capping, editing, pre-mRNA splicing, 3′ end cleavage and polyadenylation, assembly of export-competent RNPs, RNA localization, translation, stability and degradation. More than 600 RBPs are annotated in the mammalian genome based on the presence of characterized RNA binding domains, and recent experiments suggest that ∼1,000 proteins expressed by cells have RNA binding activity ( 3, 4). This process begins during transcription and is fundamental for the maturation and stabilization of RNAs ( 1, 2). 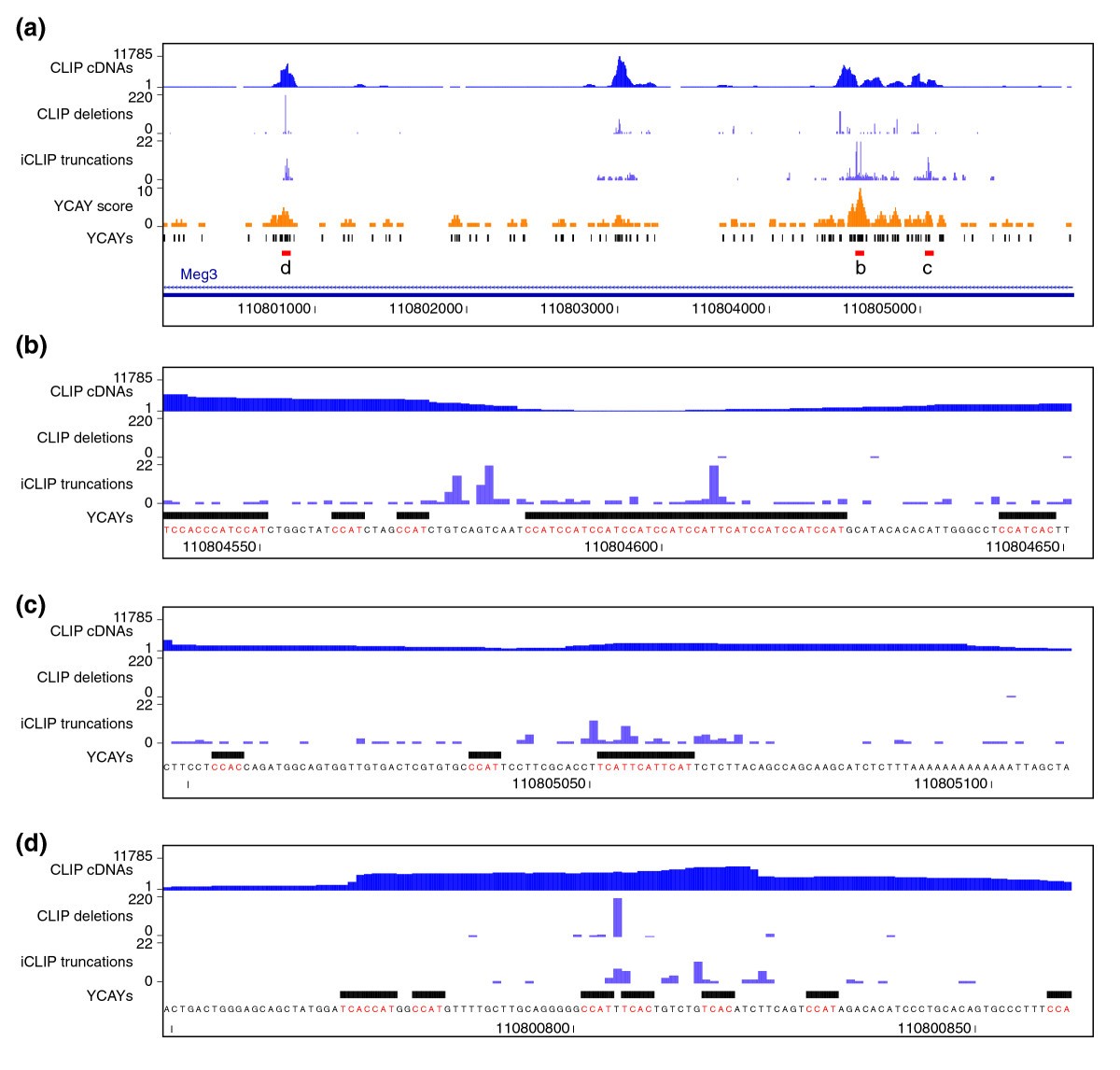
Most RNA classes are bound by RNA binding proteins (RBPs), thus forming ribonucleoproteins (RNPs). RNAs are rarely, if ever, alone in the cell. Hence, Fr-iCLIP detects the cellular location of RNA–protein interactions and provides insight into co-transcriptional, post-transcriptional and cytoplasmic RBP functions for coding and non-coding RNAs. These findings suggest that intron-retained mRNAs in the cytoplasm have previously uncharacterized functions and/or escape surveillance. However, many intron-retained mRNAs were not substrates for nonsense-mediated decay (NMD), even though they were detected in polysomes. These cytoplasmic introns were found to be highly conserved and introduced premature termination codons into coding regions. 
Surprisingly, hundreds of cytoplasmic intron targets were detected. Cytoplasmically-bound mRNAs reflected distinct functional groupings, suggesting coordinated translation regulation. As expected, SRSF3 and SRSF7 targets were detected in all fractions, with intron, snoRNA and lncRNA interactions enriched in the nucleus. To do so, we developed Fractionation iCLIP (Fr-iCLIP), in which chromatin, nucleoplasmic and cytoplasmic fractions are prepared from UV-crosslinked cells and then subjected to iCLIP. Towards this end, we sought to determine how two SR proteins-SRSF3 and SRSF7, regulators of pre-mRNA splicing, nuclear export and translation-interact with RNA in different cellular compartments. How RNA–protein interactions change over time and space to support these roles is poorly understood. 
RNA binding proteins (RBPs) regulate the lives of all RNAs from transcription, processing, and function to decay. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
